 It’s important to know what you’re eating, especially if you suffer from a food allergy.
It’s important to know what you’re eating, especially if you suffer from a food allergy.
About 220 million people worldwide live with a food allergy.1 These numbers, along with the complexity and severity of conditions, continue to rise. In America, there are about 32 million food allergy sufferers—5.6 million of those are children under the age of 18.2.2 That’s 1 out of every 13 children, or about 2 in every classroom. From a financial perspective, the cost of food allergy childcare for US families is up to $25 billion annually.
What is a food allergy?
A food allergy is an adverse health effect resulting from a specific immune response that occurs reproducibly from exposure to a given food. Food allergens are proteins that can be tolerated by most people but, in some sensitive individuals, can cause a severe, even life-threatening, reaction known as anaphylaxis.
There is no cure for severe food allergy, so complete avoidance is required. Allergic consumers rely heavily on product labeling to help them do just that. There are more than 170 foods that are reported to cause allergenic reactions. In the US, the 8 major food allergens responsible for most of the severe reactions must be declared: eggs, fish, milk, peanuts, tree nuts, shellfish or crustacean, soy and wheat.2 In the UK this list also includes 6 additional food allergens: celery, lupin, mollusks, mustard, sesame seeds and sulfur dioxide and sulfites.3
Testing is critical to ensure food safety
A study led by Michelle Colgrave and James Broadbent of the Commonwealth Scientific and Industrial Research Organization (CSIRO) found that common methods, such as the antibody-based ELISA, are not always appropriate in complex food matrices. Drawing from their experience with gluten detection using liquid chromatography-mass spectrometry (LC-MS), they developed an alternative, complementary proteomics approach to detect allergenic proteins. This approach could be the first step toward the development of a routine food testing assay.
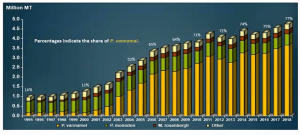
Colgrave and Broadbent’s study focused on seafood allergy for the following reasons:
- Crustacean shellfish allergies are among the most severe, with a prevalence of 2% in adults.
- It’s rare to outgrow shellfish allergy. Studies have also shown that food allergies such as shellfish are more likely to develop during adulthood.4,5
- Shrimp and prawns are 2 of the most common allergic foods. The study also looked at a mollusk (Pacific oyster) to apply the technology more broadly and to look at crossover effects
From the target groups, 3 types of shrimp and prawns were chosen based on their production worldwide. (Whiteleg shrimp is one of the most commonly caught aquatic species.)
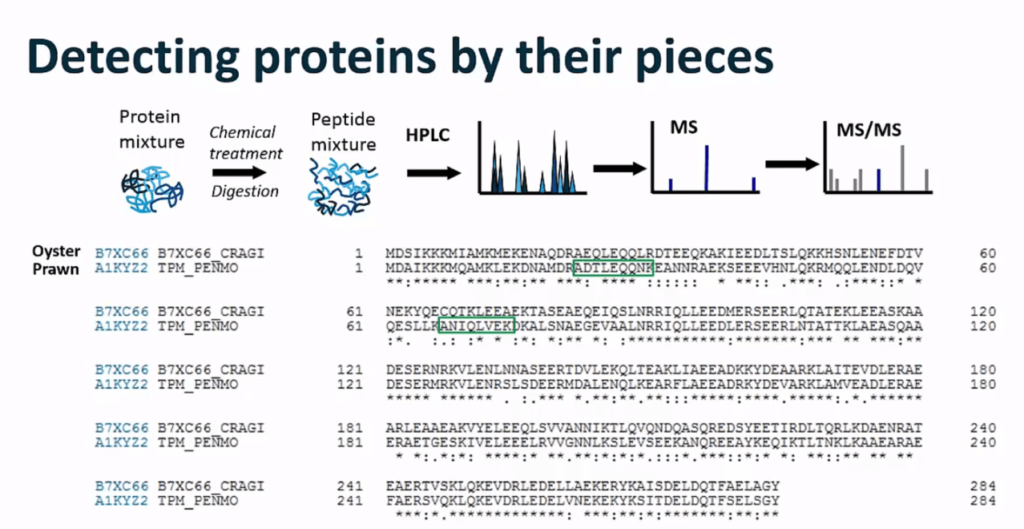
Detecting proteins by their pieces
The analysis followed these steps:
- Proteins were extracted from the food sample.
- The proteins were denatured to remove the tertiary structure. Reducing agents and alkylating agents serve to destroy disulfide bonds. These bonds hold the protein in its defined structure. By eliminating these bonds, proteins become easier to access via proteolytic enzymes.
- Enzymes (typically trypsin) were added to cut the protein into smaller pieces known as peptides. These peptides are what is detected and quantified by LC-MS.
- The peptide mixture was analyzed. The protein sequence for tropomyosin in oysters and prawns is quite different. LC-MS enabled differentiation of the peptides between the species.
The generic workflow for protein detection and quantification using LC-MS
Before you watch the webinar, here’s a summary of the research approach.
- Determine the food allergen source. (In this case, crustacean shellfish.)
- Identify potential target proteins. This study focused on tropomyosin, which was the first allergen uncovered in crustacea. It’s also responsible for food allergy in oysters. More recently, arginine kinase, myosin light, and heavy chain and sarcoplasmic calcium-binding protein have been added to the list of allergy-inducing proteins in shellfish.
- Choose a branch of the workflow. You can follow the workflow down the right (3b) of this schematic and select potential peptide markers for each protein. From their experience, obtaining experimental data by following the left (3a) path of the workflow is more likely to guide you to success. The left side is also faster.
To learn more about their work, watch their webinar by filling out the form on your right, where they describe their ongoing work on the proteome analysis of shellfish. They share data from the initial detection and identification of shellfish proteins by LC-QqTOF, and some early results of targeted allergen analysis using LC-QqQ mass spectrometry. They conclude with their goals for the second phase of the project.
Fill out the form on your right to watch the webinar.
RUO-MKT-18-10425-A
- (2017, October 25). Food Allergy Research & Education Convenes Global Leaders in Food Allergy for “Partners in Action Day” at Annual International Food Allergy & Anaphylaxis Alliance Meeting. Retrieved from https://www.foodallergy.org/about/media-press-room/food-allergy-research-education-convenes-global-leaders-in-food-allergy-for
- Facts and Statistics. Retrieved from https://www.foodallergy.org/life-with-food-allergies/food-allergy-101/facts-and-statistics
- (2018, December 21). Allergen guidance for food businesses. Retrieved from https://www.food.gov.uk/business-guidance/allergen-guidance-for-food-businesses
- Kamdar, T. A., Peterson, S., Lau, C. H., Saltoun, C. A., Gupta, R. S., & Bryce, P. J. (2015). Prevalence and characteristics of adult-onset food allergy. The Journal of Allergy and Clinical Immunology: In Practice, 3(1). doi: 10.1016/j.jaip.2014.07.007
- Sicherer, S. H., Muñoz-Furlong, A., & Sampson, H. A. (2004). Prevalence of seafood allergy in the United States determined by a random telephone survey. Journal of Allergy and Clinical Immunology, 114(1), 159–165. doi: 10.1016/j.jaci.2004.04.018






 Contact Support
Contact Support
0 Comments