Abstract
This technical note highlights the power of electron activated dissociation (EAD) for clear differentiation of 3- and 4- hydroxyproline (Hyp) isomers to meet the analytical need for biotherapeutic characterization in a case study. The elucidation of Hyp isomers poses an analytical challenge to traditional chromatographic and tandem mass spectrometry (MS/MS) approaches. By contrast, EAD generates signature z−27 and z−43 fragments for 3- and 4-Hyp isomers, respectively, for the unambiguous differentiation of these 2 Hyp isomers.
Natural linker peptides containing proline are commonly used in biotherapeutics.1 These peptides can undergo structural changes, such as Hyp formation and isomerization, during the manufacturing process.2,3 Therefore, there is a need to identify and differentiate Hyp isomers in the development of biotherapeutics to ensure their quality. However, the differentiation of Hyp isomers is challenging using traditional collision-based MS/MS approaches.3 Although conventional electron-based MS/MS techniques can generate signature fragments for the differentiation of Hyp isomers,3 these methods are not suitable for routine biotherapeutic characterization due to slow scan rate and low sensitivity. EAD has proven to be an indispensable tool for rapid differentiation of amino acid isomers, primarily when baseline chromatographic separation cannot be achieved.4-6
In this technical note, optimized EAD workflows were employed to differentiate 3- and 4-Hyp isomers of proline-containing peptides using the ZenoTOF 7600 system equipped with the EAD capability. The detection of diagnostic fragments z−27 for 3-Hyp and z−43 for 4-Hyp enabled confident elucidation of synthetic Hyp-containing peptides and a Hyp isomer identified in real-world samples from the biotherapeutic industry.
Key features of EAD for isomer differentiation and biotherapeutic characterization
- Unambiguous differentiation of isomers: EAD generates signature fragments to distinguish amino acid isomers confidently.
- High-quality MS/MS data: Gain a 5- to 10-fold increase in MS/MS sensitivity or confident isomer differentiation.
- Accurate localization: EAD preserves labile post-translational modifications (PTMs) for their accurate localization.
- Fast and flexible: EAD operates in fast and flexible modes of data-dependent acquisition (DDA) or MRMHR with a rapid scanning rate (~20 Hz in DDA mode) and the ability to tune the electron kinetic energy (KE).
- Single-injection method: The streamlined EAD DDA workflow provides comprehensive characterization of biotherapeutics in a single injection.
Introduction
The isomerization of amino acid residues, such as aspartic acid (Asp) vs. isoaspartic acid (isoAsp), leucine (Leu) vs. isoleucine (Ile) and 3-Hyp vs. 4-Hyp, can significantly impact the structure and function of biotherapeutics.7-9 Therefore, the differentiation of amino acid isomers plays a role in achieving full characterization of a therapeutic molecule at various stages of its development. It is challenging to differentiate amino acid isomers using conventional collision-based MS/MS. In this study, streamlined EAD workflows were employed to differentiate synthetic 3- and 4- Hyp isomers and determine a Hyp isomer identified in tryptic digests of a reagent protein.
Methods
Samples preparation: Two synthetic bradykinin peptides RPXGFSPFR (X = 3-Hyp or 4-Hyp) were obtained from AnaSpec Inc. The G4P linker peptides containing 3- or 4-Hyp were synthesized and purified at an Amgen facility. Synthetic peptide samples were reconstituted to a final concentration of 10 µM. The digest of a reagent protein was prepared using trypsin. 1-5 µl of each sample was injected for LC-MS analysis.
Chromatography: Synthetic bradykinin and G4P linker peptides were separated using the LC gradients 1A and 1B, respectively (Table 1), while the reagent protein digest was analyzed with the LC gradient 2 (Table 1). Peptide separation was performed using a Waters ACQUITY CSH C18 column (2.1 mm x 50 mm, 1.7 µm, 130 Å, Waters). The flow rate was set to 0.25 mL/min for synthetic Hyp-containing peptides and 0.45 mL/min for the reagent protein digest. The column was kept at 60°C in the column oven of an ExionLC AD system (SCIEX). The mobile phases A and B consisted of 0.1% formic acid (FA) in water and 0.1% FA in acetonitrile, respectively.
Differentiation of Hyp isomers by EAD
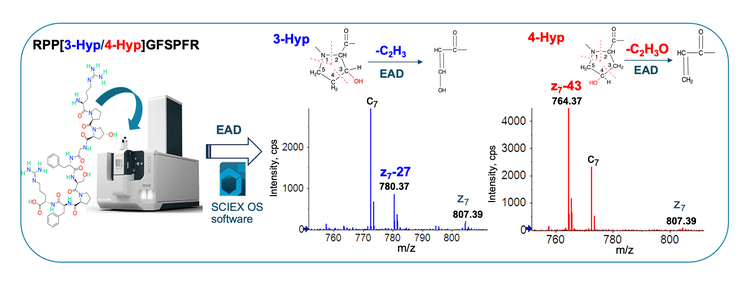
Hydroxylation of a proline residue in CHO-expressed proteins can lead to the formation of 3- and 4-Hyp isomers. The differentiation of these 2 Hyp isomers provides insight into the effect of this modification on the therapeutic molecule. Traditional collision-based MS/MS approaches do not generate diagnostic fragments to differentiate 3- and 4-Hyp isomers.3 By contrast, EAD provides a clear differentiation of Hyp isomers by producing signature z−27 and z−43 fragments for 3- and 4-Hyp, respectively.
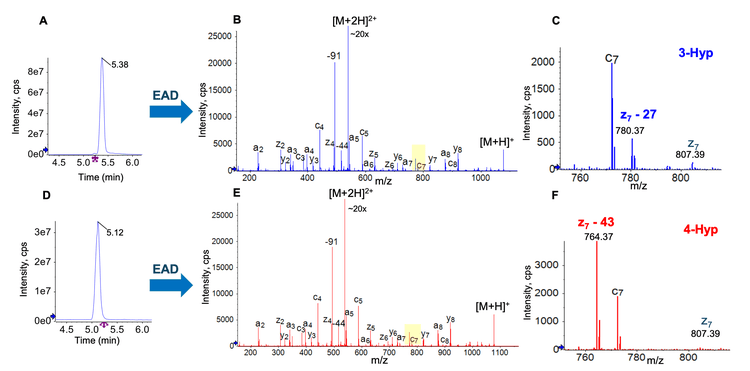
In this work, EAD was first employed to characterize 2 synthetic Hyp-containing peptides RPXGPSPFR (X = 3- or 4-Hyp). These 2 peptides were eluting at similar retention times (RTs) using the gradient 1A shown in Table 1 (Figure 2A and 2D). EAD led to rich MS/MS spectra containing Hyp-signature fragments for confident sequence confirmation and unambiguous differentiation of Hyp isomers (Figure 2B and 2E). The zoomed-in views of EAD spectra showed the diagnostic ions of z7−27 for 3-Hyp (Figure 2C) and z7−43 for 4-Hyp (Figure 2F). The neutral losses of 27 Da and 43 Da correspond to the cleavage of C2H3 and C2H3O from the pyrrolidine rings of 3- and 4-Hyp, respectively (Figure 1). The detection of these diagnostic fragments enabled confident differentiation of 2 Hyp isomers.
Differentiation of Hyp isomers of the G4P linker peptides by EAD
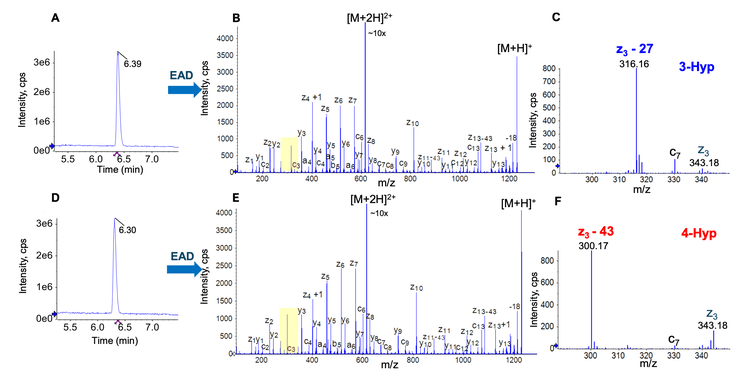
The proline residue in the linker peptide―GGGGP―might be hydroxylated and converted into 3- or 4-Hyp during the manufacturing process. To further highlight the power of EAD for Hyp isomer differentiation, 2 G4P linker peptides (SLSLSPGGGGGXAR where X = 3- or 4-Hyp) were synthesized and characterized by EAD. Similar to the example described above (Figure 2), the 2 Hyp-containing G4P linker peptides were eluting at close RTs (Figure 3A and 3D). Excellent EAD fragmentation of 2 isomers led to the detection of a nearly complete series of c/z fragments (Figure 3B and 3E). The expanded views of EAD spectra showed the detection of a z3−27 fragment at m/z 316.16 for 3-Hyp (Figure 3C) and a z3−43 ion at m/z 300.17 for 4-Hyp (Figure 3F). These results further demonstrate the power of EAD for precise and rapid differentiation of 3- and 4-Hyp isomers of the proline-containing linker peptides.
Case Study: Elucidation of a Hyp isomer identified in real-world IL-2 samples
E. coli expressed reagent proteins have been approved as an oncology therapeutic. However, this version of reagent protein has a short in vivo half-life and toxic side effects when administered at high doses. Hence, there is considerable interest in developing CHO-expressed reagent protein. Recent LC-MS analyses of real-world tryptic digests from several CHO-expressed proteins led to the unexpected identification of a Hyp-containing peptide (data not shown). EAD was leveraged to determine whether hydroxylation of the proline residue occurred in the 3rd or 4th position of the pyrrolidine ring.
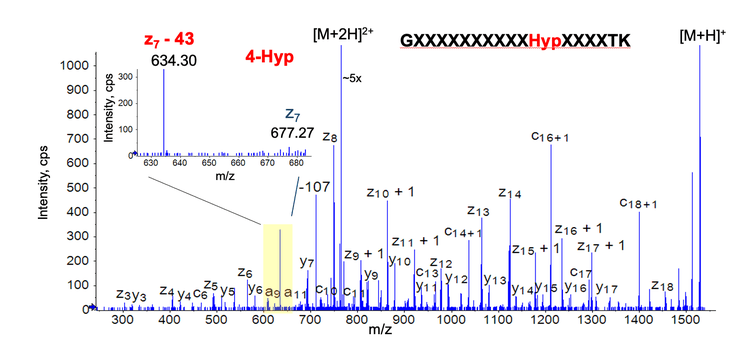
Figure 4 shows the EAD spectrum of the doubly charged peptide GXXXXXXXXXXHypXXXXTK from the reagent protein (amino acid residues marked “X” were covered for proprietary reasons). EAD generated rich sequence fragments for confident identification of this Hyp-containing peptide in the reagent protein digest. The detection of a diagnostic ion z7−43 (loss of C2H3O) at m/z 634.3 confirmed the presence of 4-Hyp instead of 3-Hyp in this reagent protein peptide (inset in Figure 4).
In summary, the data presented in this technical note demonstrates the power of EAD for confident and rapid elucidation of Hyp-containing peptides that are challenging to characterize using traditional chromatographic or fragmentation techniques. The ability of EAD for confident isomer differentiation was successfully leveraged to determine a Hyp isomer identified in real-world samples from the biotherapeutic industry.
Conclusion
- EAD provided confident identification and clear differentiation of 3- and 4-Hyp isomers that are challenging to elucidate using traditional fragmentation techniques.
- EAD produced the signature z−27 and z−43 fragments for 3- and 4-Hyp isomers, respectively, enabling unambiguous differentiation of these 2 isomers.
- Streamlined EAD methods were employed to differentiate synthetic peptides containing 3- and 4-Hyp residues confidently.
- An unexpected Hyp-containing peptide identified in the tryptic digests of a reagent protein was the 4-Hyp isomer based on the detection of a signature z−43 fragment using EAD.
- EAD workflows described in this technical note can be leveraged to identify and differentiate Hyp-containing peptides from protein therapeutics rapidly.
References
- Arnik Shah et al. (2024) Characterization of charge variants, including post-translational modifications and proteoforms, of bispecific antigen-binding protein by cation-exchange chromatography coupled to native mass spectrometry. Talanta. 266(1):125062.
- Oksana Tyshchuk et al. (2019) Characterization and prediction of positional 4-hydroxyproline and sulfotyrosine, two post-translational modifications that can occur at substantial levels in CHO cells-expressed biotherapeutics. MAbs. 11(7):1219-1232.
- Fengfei Ma et al. (2018) Large-scale differentiation and site specific discrimination of hydroxyproline isomers by electron transfer/higher-energy collision dissociation (EThcD) mass spectrometry. Anal. Chem. 90(9):5857-5864.
- Takashi Baba et al. (2021) Dissociation of Biomolecules by an Intense Low-Energy Electron Beam in a High Sensitivity Time-of-Flight Mass Spectrometer. J. Am. Soc. Mass Spectrom. 32(8):1964-1975.
- Comprehensive differentiation isomers from forced degradation by electron activation dissociation (EAD). SCIEX technical note, RUO-MKT-02-14730-A.
- An evaluation of single injection platform method for advanced characterization of protein therapeutics using electron activation dissociation (EAD). SCIEX technical note, RUO-MKT-02-13965-A.
- Mary Krause and Erinc Sahin. (2019) Chemical and physical instabilities in manufacturing and storage of therapeutic proteins. Current Opinion in Biotechnology. 60:159-167.
- Anshu Kuriakose et al. (2016) Immunogenicity of Biotherapeutics: Causes and Association with Posttranslational Modifications. J. Immunol. Res. 2016:1298473.
- Anna Robotham and John Kelly (2020) LC-MS characterization of antibody-based therapeutics: recent highlights and future prospects. Approaches to the Purification, Analysis and Characterization of Antibody-Based Therapeutics. Chapter 1:1-33.


