Structural characterization of the cationic lipid nanoparticle component, ALC-0315, and its impurities using electronactivated dissociation (EAD)-based MS/MS fragmentation
Featuring the ZenoTOF 7600 system
Zhichang Yang1 , Paul Baker1 , Sahana Mollah1 , Robert Proos1 and Jonathan Le Huray2
1SCIEX, USA; 2Acuitas Therapeutics Inc
Abstract
In this technical note, [(4-hydroxybutyl)azanediyl]di(hexane-6,1- diyl) bis(2-hexyldecanoate), commonly known as ALC-0315, and its impurities at relative abundances as low as 0.19% were structurally characterized using EAD fragmentation on the ZenoTOF 7600 system. Collision-induced dissociation (CID) did not generate sufficient fragments to fully characterize ALC-0315 and impurities. In contrast, EAD provided abundant diagnostic fragments to allow near-complete structural elucidation of singly charged compounds (Figure 1).
Introduction
The use of lipid nanoparticles (LNP) as drug delivery devices has dramatically increased since the advent of the COVID-19 vaccine and introduction of recent gene therapy therapeutics. Lipid impurities in the LNP can attenuate the effectiveness of the active pharmaceutical ingredient (API). For example, N-oxidation of ionizable lipids might lead to covalent modification of ribonucleotides and a loss of mRNA potency. 1 It is therefore necessary to ensure detailed and sensitive characterization of the ionizable lipid and its related impurities. However, detailed characterization of LNPs is challenging using current liquid chromatography-mass spectrometry (LC-MS) methodologies. EAD can be used to provide structurally diagnostic fragment ions to elucidate the distinctive structures of complex lipids. 2-8
Here, the applicability of EAD for the detailed characterization of ALC-0315 and its impurities was tested. This lipid plays a crucial role in the function of LNPs by stabilizing the negatively charged mRNA and its impurities might affect the therapeutic effectiveness of the LNP formulation. 8 Using the ZenoTOF 7600 system equipped with EAD, ALC-0315 and its raw materialrelated impurities were fully characterized. The nature and site of chemical alterations in ALC-0315 resulting from forced degradation were also elucidated using structurally diagnostic fragment ions produced by EAD.
Figure 1. MS/MS spectra acquired for ALC-0315 (m/z = 766.7) using EAD (left) and CID (right). Blue peaks were assigned to a fragment of the structure. Red peaks were not assigned. More than 70% of the EAD-generated peaks were assigned to fragments. EAD generated many more assigned fragments than CID, including an important fragment from the head group of the molecule (m/z = 172) that was used for structural elucidation.
Key features of the comprehensive LNP characterization
- EAD fragmentation using the ZenoTOF 7600 system provided informative fragment ions to structurally characterize ALC-0315 with minimal tuning
- Information-rich MS/MS spectra were used for definitive and distinct structural elucidation of low-abundant impurities, including oxidation and hydroxyl-to-carboxyl transformation
- Detection and identification of lipid impurities at levels <0.2% abundance relative to the main peak were achieved using the Zeno trap
Methods
Chemicals and materials: ALC-0315 was provided by Acuitas Therapeutics Inc. Methanol (MeOH) was purchased from LiChrosolv (PN: 1.06035.2500). Ammonium acetate was from Fluka Analytical (PN: 73594-25G). Acetonitrile was (ACN) from Sigma-Aldrich (PN: 34851-4L).
Sample preparation: ALC-0315 standard suspended in PBS with 300mM sucrose was aliquoted 5 mg per vial and subjected to 60°C incubation for 3 days or 5 days for forced degradation.
The intact and degraded ALC-0315 stock solutions were diluted 100-fold with a solution of 60:40 (v/v), ACN/MeOH with 10 mM ammonium acetate (mobile phase B). The resulting solution was further diluted 100-fold with a solution containing 15% water, 30% MeOH and 55% ACN with 10mM ammonium acetate (mobile phase A). The solution was then analyzed directly by
LCMS/MS. LC-MS/MS analysis: Samples were analyzed by a LC-MS/MS system equipped with a Waters H-class UPLC and a SCIEX ZenoTOF 7600 system. The samples were separated using a UHPLC Peptide BEH C18 column (2.1 × 150 mm, 1.7 μm, 300 Å, Waters, PN: 186003687) using the gradient conditions shown in Table 1. The flow rate was set to 0.3 mL/min and the column temperature was set to 70°C. The injection volume was set to 2 µL.
Table 1. LC gradient.
LC-MS/MS data were acquired using a data-dependent acquisition (DDA) scan mode. The parameters used in these experiments are shown in Tables 2 and 3.
Table 2. TOF MS parameters.
Table 3. TOF MS/MS parameters.
Data processing: Structural elucidation was performed and relative quantification was determined using the Explorer module of SCIEX OS software.
Figure 2. TIC of ALC-0315 exposed to forced degradation. The structure of ALC-0315 is shown in the yellow inset square. Forced degradation of ALC-0315 was achieved by incubating the compound at 60°C for 3 or 5 days. The m/z values for 4 impurities are shown above the TIC. These degradation products included the loss of an acyl chain (m/z = 428.4087), the loss of the head group (m/z = 694.6690), oxidation (m/z = 782.7216) and the loss of an alkyl chain (m/z = 708.6495).
Forced degradation yields increased in impurities
Forced degradation experiments were performed to expedite the formation of impurities. Impurities might be formed during the LNP formulation process because it involves heat and mechanical stress. Therefore, it is important to identify and report any impurities that are formed.
Figure 2 shows the total ion chromatogram (TIC) for the differentially treated samples. The intensities of impurities, represented by the smaller peaks in the TIC, appeared or increased during the incubation period. The m/z of each impurity and ALC-0315 is shown above the TIC. Based on the formula finder module in SCIEX OS software, the peak at m/z 428.4087 likely corresponds to a loss of the acyl chain from the intact ALC0315. Similarly, the peak at m/z 694.6690 likely corresponds to the loss of the head group, the peak at m/z 782.7216 likely corresponds to the addition of O and the peak at m/z 708.6495 likely corresponds to the loss of an alkyl chain (C4H9) and desaturation.To confirm the structure of these impurities, MS/MS interpretation is necessary.
The 5-day incubation time was expected to provide the greatest distribution of ALC-0315 impurities and therefore the data from this sample were analyzed in detail. Figure 3 shows the extracted ion chromatogram (XIC) of ALC-0315 and several lowabundant impurities. ALC-0315 eluted at 12.06 min. The impurity peaks were observed at intensities ranging from 5.9% to 0.2% relative to ALC-0315. These results demonstrate the wide interscan dynamic range of the ZenoTOF 7600 system for lipid impurity analysis. Table 4 summarizes the various impurities identified and their intensities relative to the ALC-0315 peak based on the TOF MS peak area.
Table 4. Compounds identified by EAD in ALC-0315-5D
Figure 3. XIC of intact ALC-0315 and 6 low-abundant impurities after 5 days of forced degradation. The top panel shows the normalized XIC of all compounds, in which ALC-0315 (peak 1) is the base peak of the chromatogram. The bottom panel shows the same chromatogram with a 10x zoom of the y-axis to better visualize the low-abundant impurities (peaks 2-7). Impurities were identified at relative intensities as low as 0.2%.
Structural elucidation of ALC-0315 and selected impurities
MS/MS data enabled the detailed structural elucidation of ALC0315 and impurities. Structural information for peaks 1, 2 and 7 (Figure 3) is presented here. The characterization of peaks 3, 4, 5 and 6 is included in the supplemental information.
The m/z observed by TOF MS for the singly charged ALC-0315 precursor ion (Figure 3, peak 1) matched the theoretical m/z of ALC-0315 within 2 ppm (Table 4). This ion was selected for fragmentation by EAD (Figure 1, left) or CID (Figure 1, right). EAD provided a more information-rich spectrum than CID. More than 100 peaks were assigned to structures from the in silicoderived fragment library with <0.01 Da mass error. In contrast, CID generated approximately 10 fragment ions. The relatively high signal of the peak at m/z 528.5042 (Figure 1, left) indicates that EAD preferentially breaks the ester bond between the O and the C=O of the ester bond, whereas CID favors breaking the bond between the C and O adjacent to the ester bond. The intense peak at m/z 172.1725 (Figure 1, left) shows fragmentation of the head group of the molecule, which was not observed by CID. This fragment was important to identify the oxidation site in ALC-0315 impurities.
The peak at m/z 782.7271 (Figure 3, peak 2) was the primary impurity of ALC-0315 and occurred with 5.9% relative intensity. The m/z of this impurity was 15.9943 Da more than that of intact ALC-0315 (m/z = 766.7274), indicating that O was incorporated.
The intensity of this peak increased with the forced degradation timecourse (Figure 2, right). Figure 4 shows the EAD fragmentation spectra for ALC-0315 and the unknown oxidation product. The peak at m/z 544.4956 (Figure 4, bottom) from oxidized ALC-0315 is 15.9914 Da more than the m/z 528.5042 (Figure 4, top) fragment from intact ALC-0315. This observation indicates that O was incorporated to the left of the ester bond. Similarly, the peak at m/z 443.3988 (Figure 4, bottom) from the oxidized ALC-0315 is 15.9917 Da more than the peak at m/z 427.4071(Figure 4, top) from the intact ALC-0315, which indicates the addition of that O was added to the left of the C-N bond. Finally, the addition of 14.9854 Da (187.1579 (Figure 4, bottom)-172.1725 (Figure 4, top) = 14.9854) corresponds to the gain of one O and loss of H on the head group N atom.
Figure 4. MS/MS spectra acquired for ALC-0315 (m/z = 766.7) and its oxidized impurity (m/z = 782.7) using EAD. Blue peaks were assigned to a fragment of the structure. Red peaks were not assigned. More than 70% of the EAD-based fragment peaks were assigned for both spectra.
Figure 5 shows the CID spectra generated for ALC-0315 (top panel) and the oxidized impurity (bottom panel). The 16 Da difference between peaks at m/z 438 and 454 and peaks at m/z 694 and 710 indicates O incorporation in the oxidized ALC-0315 to the left of the ester bond. However, the diagnostic fragments containing the head group were not observed in the CID spectra to further localize the oxidation. Only EAD-based fragmentation generated the diagnostic fragments necessary for such confident structural elucidation.
Figure 5. CID-based MS/MS spectra acquired from ALC-0315 (top panel, m/z = 766.7) and its oxidized impurities (bottom panel, m/z = 782.7). Blue peaks were assigned to a fragment of the structure. Red peaks were not assigned. The lack of head group fragments prevented the localization of O incorporation. The CID data indicate that the O incorporation can be anywhere to the left of the ester bond.
EAD spectra were also used to characterize the peak at m/z 780.7061 that eluted at 10.77 min (Figure 3, peak 7). The intensity of this peak increased over the 5-day forced degradation timecourse (data not shown) and was present at 0.19% relative intensity in ALC-0315-5D sample (Table 4). Despite the relatively low abundance of this impurity peak, an information-rich EAD spectrum was successfully acquired. The intact mass of 780.7061 Da corresponds to the gain O and loss of 2 H from intact ALC-0315. Two structures are possible with this specific mass addition: 1) oxidation at the head group N and double bond formation on the alkyl chain and 2) oxidation of the hydroxyl moiety to yield a carboxylic acid. Figure 6 shows the MS/MS spectra acquired via EAD (top panel) and CID (bottom panel). The EAD spectrum confirms that the peak at m/z 780.7061 corresponds to oxidation of the hydroxyl functional group. The peak at m/z 542.4767 designates the gain of O and loss of 2 H to the left of the ester bond and the fragment at m/z 438.4278 indicates that no O is present on the N of the head group. Combined with the peak at m/z 186.1488, these data indicate that O is incorporated at and the 2 H are lost from the head group, resulting in the oxidation of the hydroxyl moiety to a carboxylic acid functional group. In contrast, the CID data (Figure 6, bottom panel) were insufficient to distinguish between the 2 possible structures.
Figure 6. MS/MS spectra acquired for the hydroxyl-to-carboxyl impurity using EAD (top panel) and CID (bottom panel) (m/z = 780.7). Blue peaks were assigned to a fragment of the structure. Red peaks were not assigned. EAD generated a fragment containing the head group, indicating the oxidation of the hydroxyl moiety to a carboxylic acid functional group.
The data generated by EAD provided better descriptions of important structural features than the same experimental data acquired using CID. CID yielded low fragmentation efficiency and most fragment ions had less than 5% relative intensity compared to the parent ion. This workflow leveraged the speed, sensitivity and broad dynamic range of the ZenoTOF 7600 system. The unique hardware configuration of the ZenoTOF 7600 system made it well-suited for the analysis and quality control of LNP formulations and their individual components. Data generated using this workflow can be used to determine the efficacy and safety of formulated LNPs and to aid the rational design of new synthetic lipids.
Conclusion
- Untargeted analysis by mass spectrometry using EAD provided an in-depth structural characterization of singly charged, ionizable lipids and related impurities with a single LC-MS/MS run
- The ZenoTOF 7600 system provides improved risk assessment of formulated LNPs through explicit structural elucidation and localization of O incorporation into impurities derived from cationic lipids such as ALC-0315
- The interscan dynamic range and quantitative sensitivity of the ZenoTOF 7600 system can decrease the risk of missing critical low-abundance impurities
References
- Jones JW, et al. Electron-induced dissociation (EID) for structure characterization of glycerophosphatidylcholine: determination of double-bond positions and localization of acyl chains. J Mass Spectrom. 2015 Dec;50(12):1327-39. PMID: 26634966
- Baba T, et al. Structural identification of triacylglycerol isomers using electron impact excitation of ions from organics (EIEIO). J Lipid Res. 2016 Nov;57(11):2015-2027. PMID: 27457033
- Baba T, et al. In-depth sphingomyelin characterization using electron impact excitation of ions from organics and mass spectrometry. J Lipid Res. 2016 May;57(5):858-67. PMID: 27005317
- Campbell JL, Baba T. Near-complete structural characterization of phosphatidylcholines using electron impact excitation of ions from organics. Anal Chem. 2015 Jun 2;87(11):5837-45. PMID: 25955306.
- Baba T, et al. Quantitative structural multiclass lipidomics using differential mobility: electron impact excitation of ions from organics (EIEIO) mass spectrometry. J Lipid Res. 2018 May;59(5):910-919. PMID: 29540574
- Baba T, et al. Development of a Branched Radio-Frequency Ion Trap for Electron Based Dissociation and Related Applications. Mass Spectrom (Tokyo). 2017;6(1):A0058. PMID: 28630811
- Baba T, et al. Distinguishing Cis and Trans Isomers in Intact Complex Lipids Using Electron Impact Excitation of Ions from Organics Mass Spectrometry. Anal Chem. 2017 Jul 18;89(14):7307-7315. PMID: 28613874
- Packer M, et al. A novel mechanism for the loss of mRNA activity in lipid nanoparticle delivery systems. Nat Commun. 2021 Nov 22;12(1):6777. PMID: 34811367
 Click to enlarge
Click to enlarge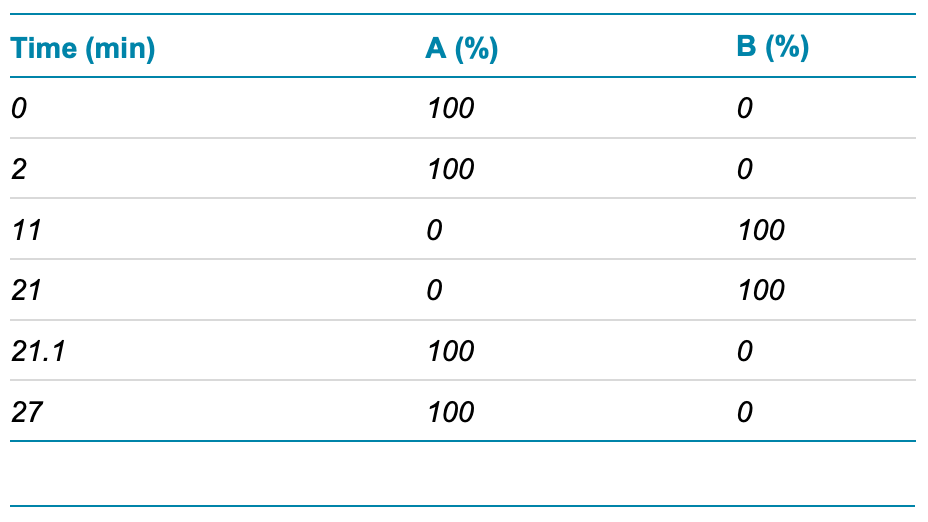 Click to enlarge
Click to enlarge Click to enlarge
Click to enlarge Click to enlarge
Click to enlarge Click to enlarge
Click to enlarge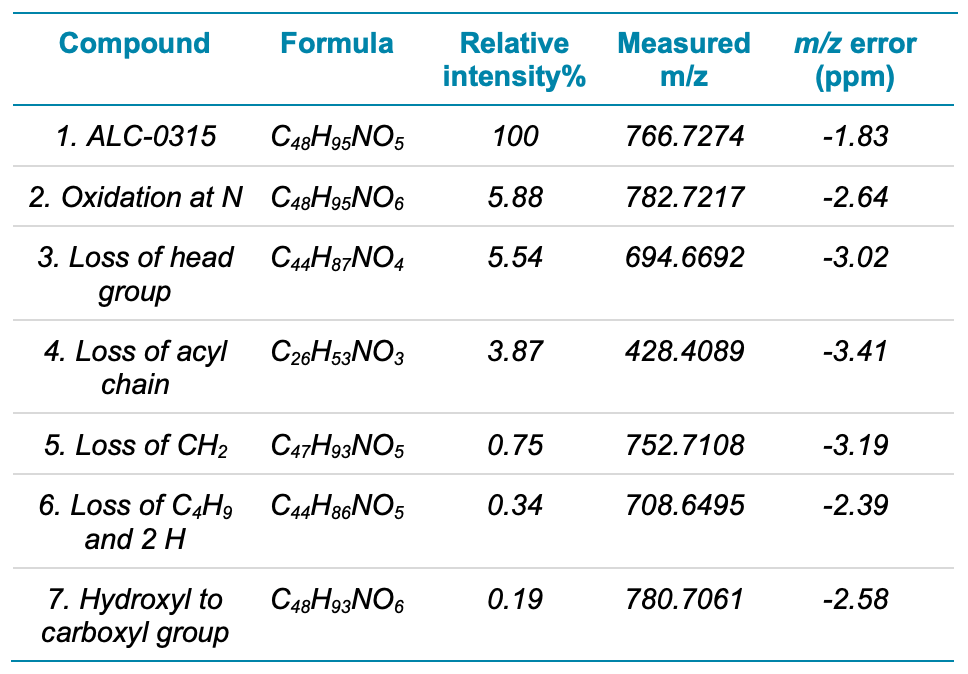 Click to enlarge
Click to enlarge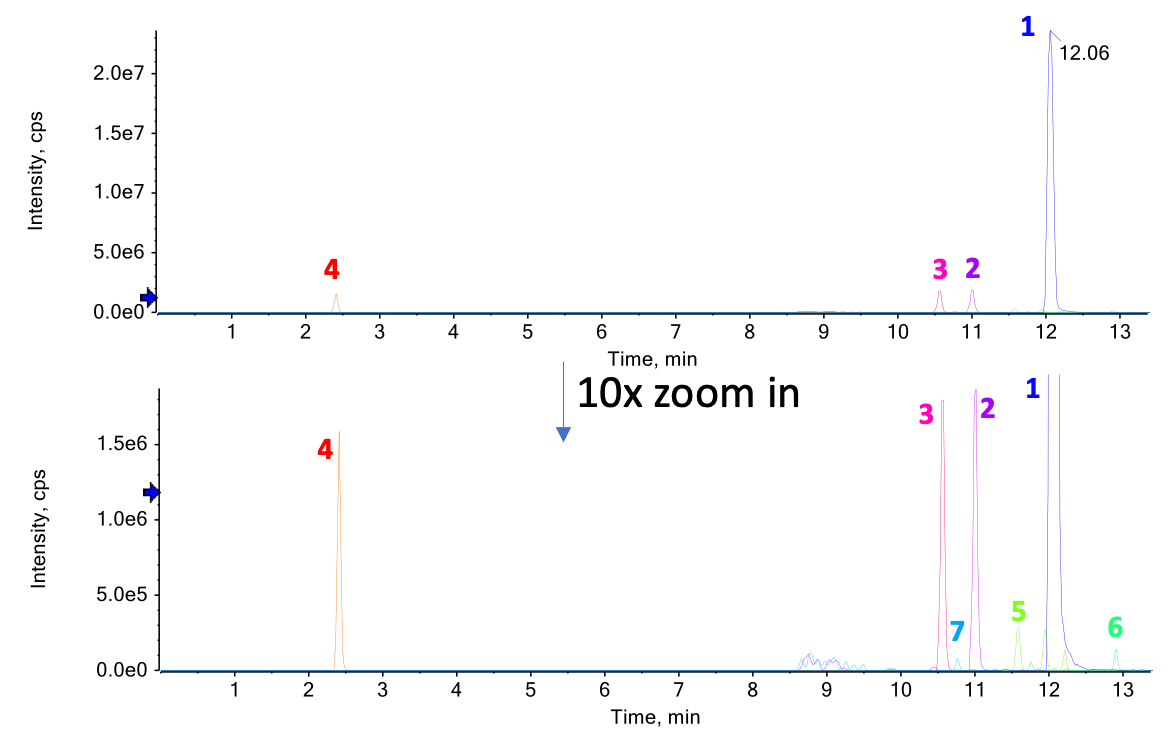 Click to enlarge
Click to enlarge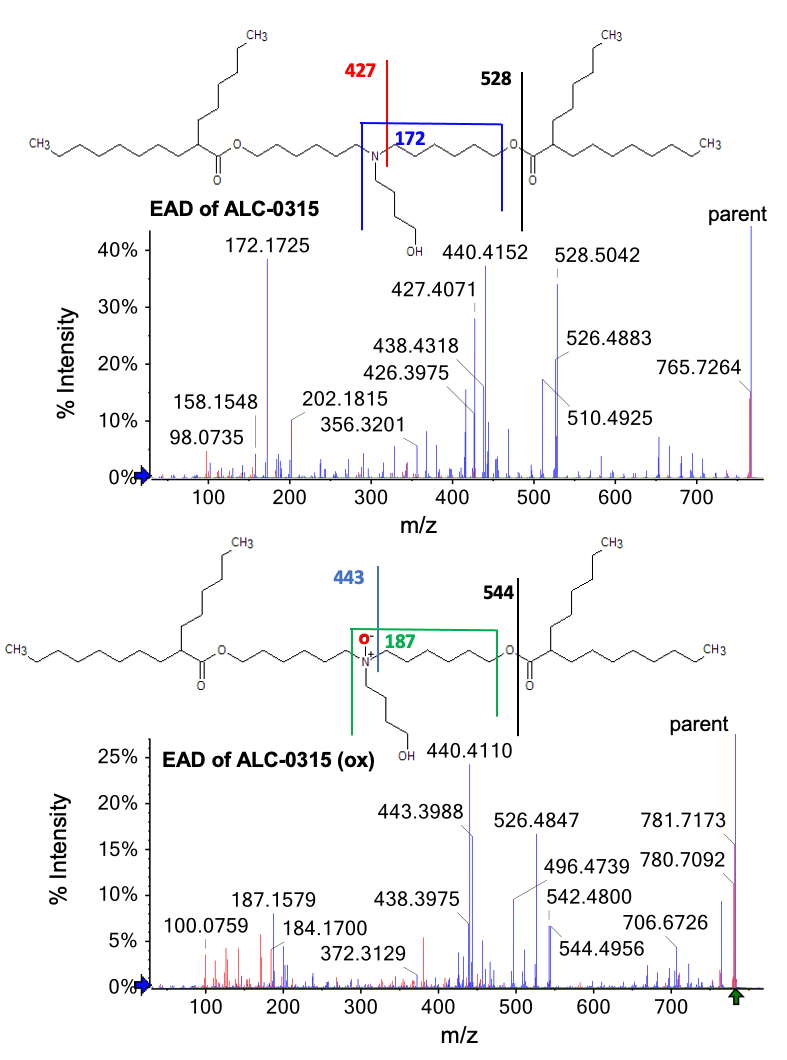 Click to enlarge
Click to enlarge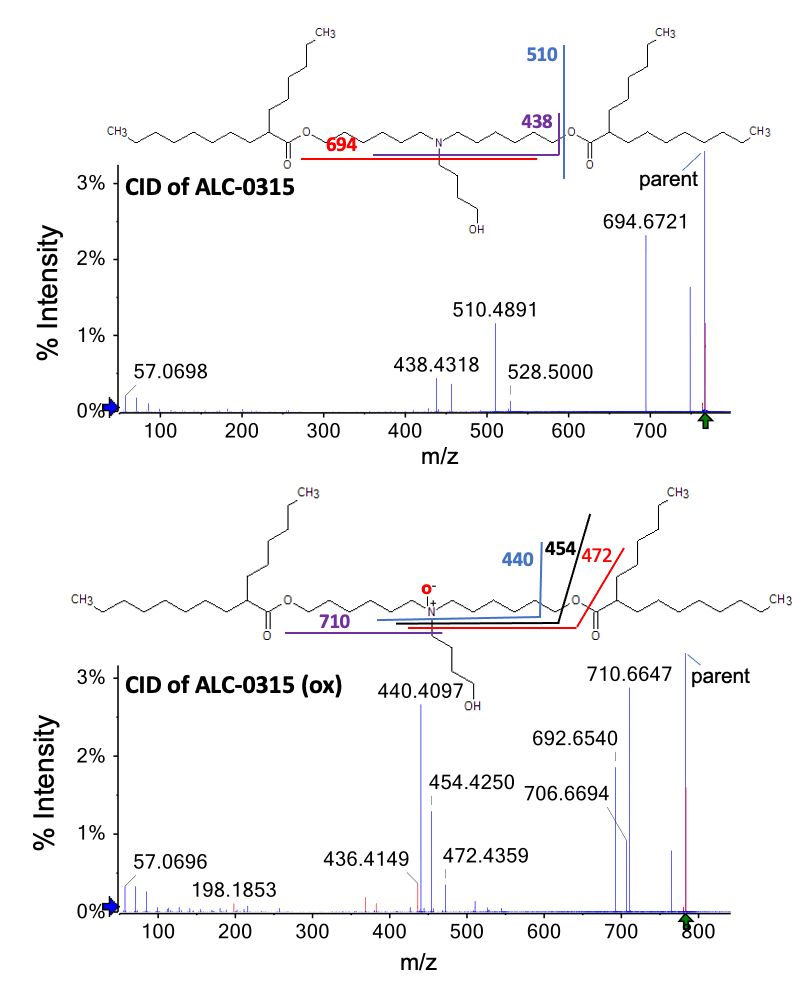 Click to enlarge
Click to enlarge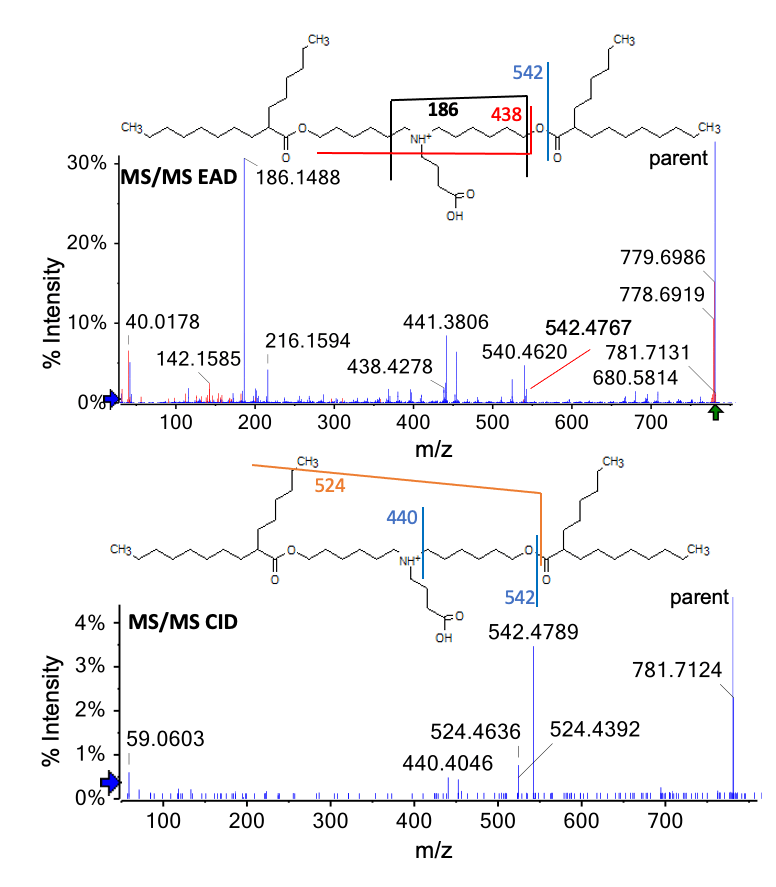 Click to enlarge
Click to enlarge