For the inquisitive data explorer, PeakView software is everything you need for spectral analysis and data interrogation. Easily review, identify, superimpose, compare samples, and label peaks in just a few clicks. PeakView software supports mass spectrometer systems for qualitative review of LC-MS and MS/MS data as well as optional detectors such as UV and DAD.

The PeakView software difference
Confidently interrogate your data
Gain rapid insight on your data. Screen and identify peaks automatically by processing LC-MS/MS data using information on retention times, accurate mass, isotopic pattern, and MS/MS library searching.
Intuitively made for you
Data review and presentation is simplified and enhanced with functions that allow comparison, identification, labeling, and creation of informative contour and overlay plots.
Flexibility so you can do great things
Interrogate data for a diversity of workflows through micro-applications including SWATH acquisition, Bio Tool Kit, and MasterView software.
Superior peak detection and deconvolution
Quick and easy qualitative review and comparison
Don’t miss anything, see everything all at once because PeakView software allows comprehensive data coverage. Identify peaks across scales and with varying intensities in a single spectrum, whether you’re a novice or advanced user. From raw mass spec data to final output result validation and visualization, informative visuals such as overlaid chromatograms or heat maps help you quickly view and identify peaks easily.
PeakView software is ideal to get an in-depth insight of your data, and the broad range of tools serves as a great first source for generating your pre-publication reports.
Sophisticated investigation and characterization at the molecular level
Quickly calculate and determine all possible elemental formulas to detect masses using both accurate mass and isotope distribution with Formula Finder. Using advanced algorithms that use chemical logic and available MS/MS data, efficiently and accurately identify and filter through non-targeted samples.
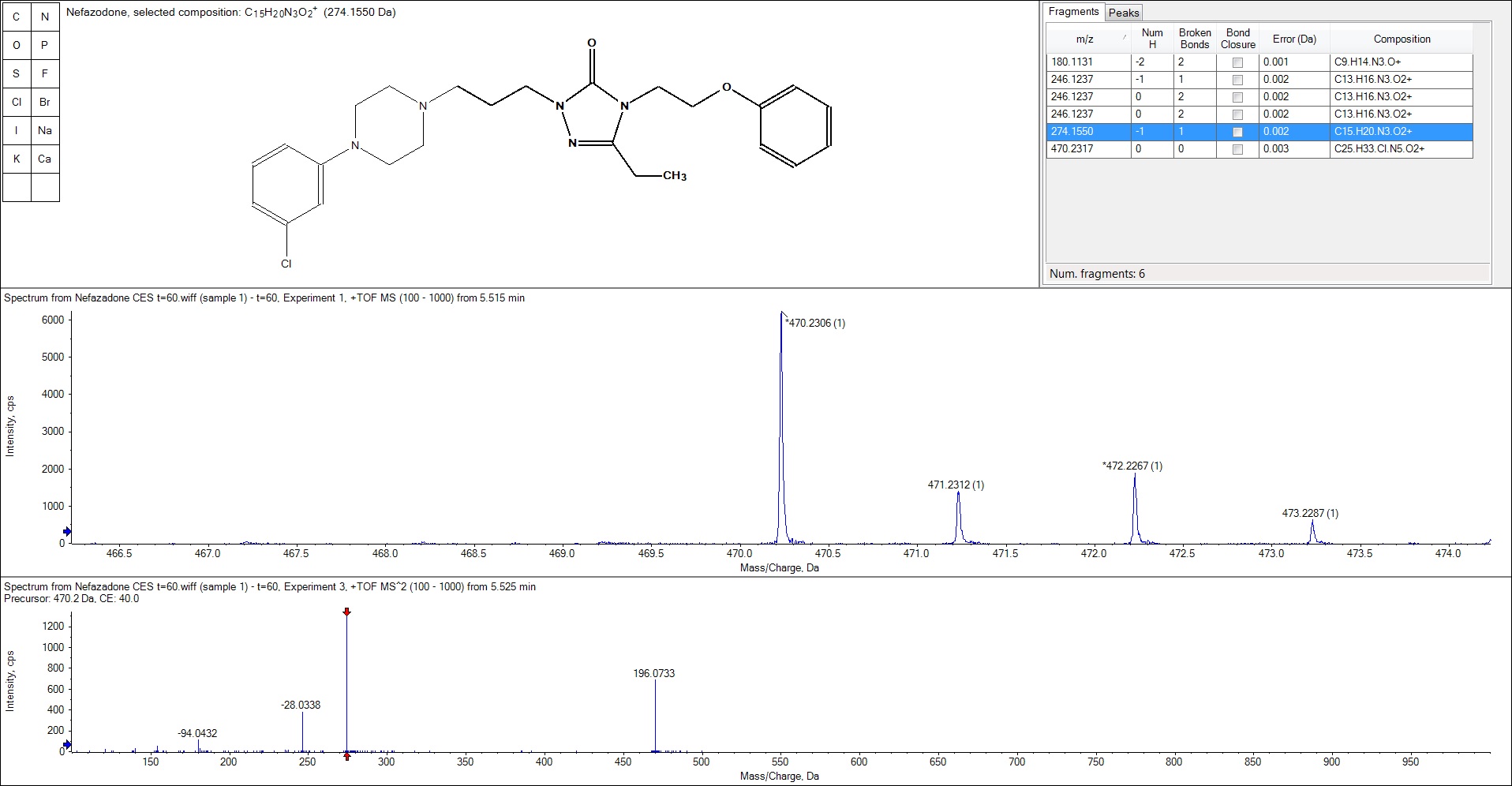
The structural elucidation in PeakView software links the masses of ions to structures to help identify and characterize compounds, explore possible sites for biotransformation, and provide insights into fragmentation mechanisms. Matching compositions and substructures to the MS and MS/MS allows users to confirm identity and explore more complex fragmentation pathways.
Integrated micro apps for enhanced functionality
Novel software micro-applications embedded into the PeakView software enhance functionality for specific workflows.
For instance, SWATH acquisition micro-application. It is a comprehensive processing tool for quantitative proteomics. Associate your protein spectral ion library with your SWATH data and easily turn the comprehensive SWATH datasets into quantitative answers. Retention time calibration, advanced extraction and scoring algorithms ensure the highest data integrity, which can then be exported to the MarkerView software for statistical analysis.
Another micro-application, Bio Tool Kit can be added to PeakView software to easily characterize your biomolecules. Whether you are characterizing your peptide by de novo sequencing, or your protein using peptide mapping and intact protein reconstruction, the intuitive, interactive tools allow you to find the right answer quickly. The extensive post-translational modification catalog ensures answers to the most challenging spectra can be determined.
Software
Resources
-
Document
PeakView 2.2 Software Release Notes
-
Document
PeakView 2.2 Software Release Notes Addendum
-
Document
PeakView software 2.2 EULA
-
Technical note
Identification of Degradation Products of Simvastatin
-
Technical note
Ultra-sensitive host cell protein detection using CESI-MS with SWATH acquisition
-
Technical note
Bio Tool Kit for Biomolecule Characterization
-
Document
Kombination von LC-QTOF-MS- und LC-Triple Quad-Messungen zum sensitiven Nachweis von synthetischen Cannabinoiden


